xarray has a built in NDPointIndex. Which is
useful for dealing with n-dimensional coordinate variables representing irregular data.
Which sounds like the same problem that NDIndex solves! However NDPointIndex and NDIndex actually solve distinct problems. and would be difficult to use to solve what is solved by the other.
NDPointIndex¶
NDPointIndex handles the case of allowing selecting by a distance metric for points distributed in N-dimensional space. For example if we data points distributed in 2 dimensions then each point will have 2 different annotaiton values associated iwth it (x, and y). This gives rise to NDPointIndex’s requirement to have the same number of coordinates as dimensions.
NDIndex¶
NDIndex solves works with annotations on top of regular gridded data. For example if we have a long 1 D dataset of continuous data collection over time (absolute time) that where every 5 seconds a new trial begins (e.g. a speech task in a neuroscience experiment) then we then break the data up into individual trials and keep track of the relative time in each trial then our data cube will have shape (trials, rel_time). If we want to keep track of the absolute time associated with each data point then that coordinate will be 2D coord (trials, rel_time). Or, more usefully we might have an event that happened at a different time in each trial (e.g. speech onset) and want to perform analysis relative to that point. For this we would nede a new coord event_locked_time which will be 2D as well.
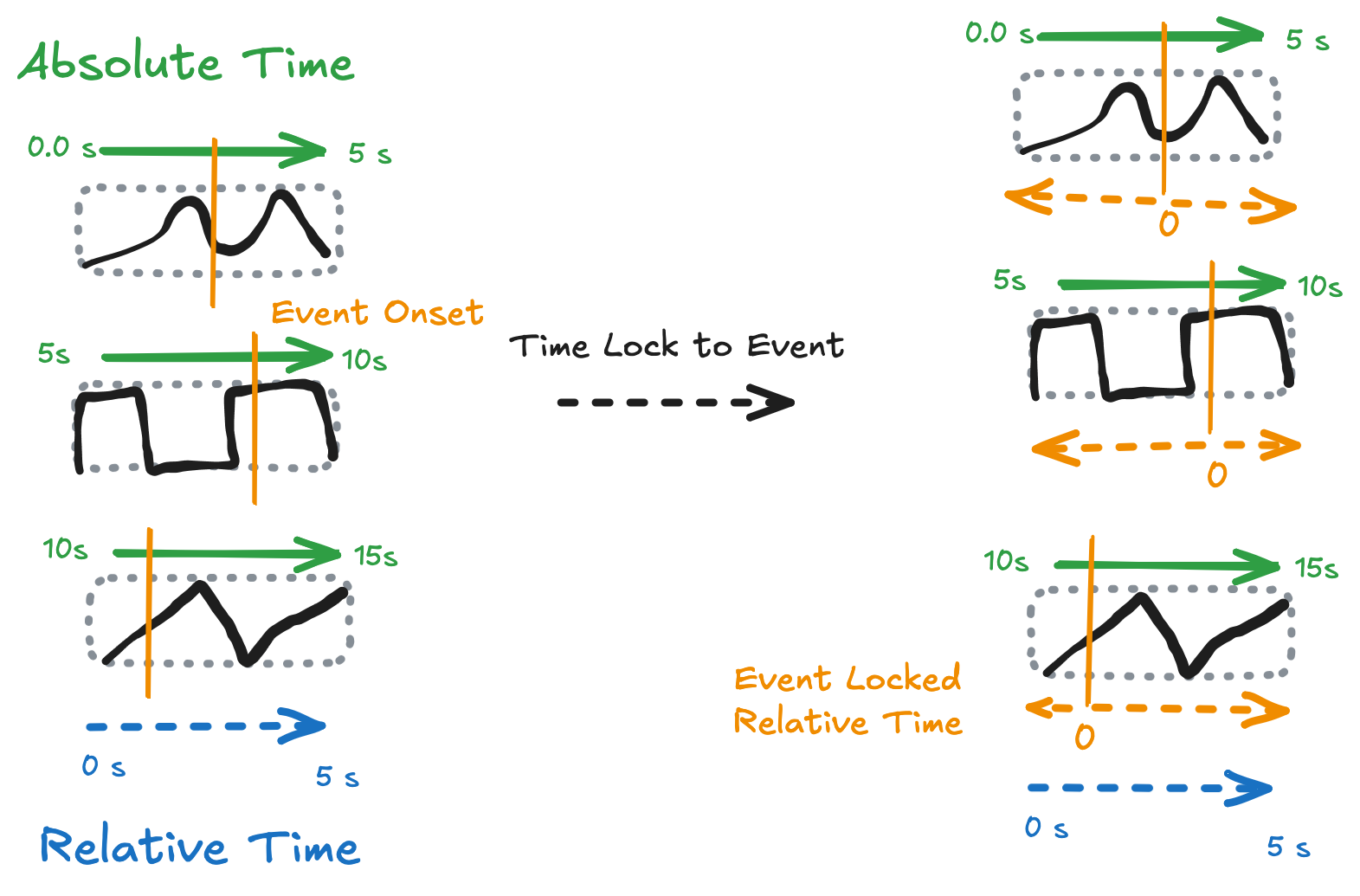
In this case we have a single annotation associated with each point in the data, even though it is 2D! i.e. each point has a unique event_locked_time. This would not be allowed by NDPointIndex
Differences¶
The fundamental difference is:
| Aspect | NDPointIndex | NDIndex |
|---|---|---|
| Coordinates | Multiple 2D coords that together define position | Single N-D coord with derived values |
| Query type | Spatial: “find point at (x, y)” | Value: “find cell where value ≈ target” |
| Use case | Curvilinear grids, scattered observations | Structured arrays with computed coordinates |
They solve different problems
NDPointIndex answers: “Which grid cell is nearest to coordinates (lat=45.2, lon=-122.5)?”
ds.sel(lat=45.2, lon=-122.5, method="nearest") # Spatial queryNDIndex answers: “Which (trial, rel_time) cells have event_time closest to .5?”
ds.sel(event_time=.5, method="nearest") # Value lookup in N-D arrayUse NDPointIndex when your coordinates define positions in space (or similar multi-dimensional coordinate systems).
Use NDIndex when you have derived coordinates computed from dimension coordinates (like event_time = rel_time - event_onset).
%xmode minimal
import numpy as np
import xarray as xr
from linked_indices.example_data import trial_based_datasetException reporting mode: Minimal
Trying NDPointIndex with trial-based data¶
Now let’s see what happens when we try to use NDPointIndex with our trial-based dataset where we have a single 2D abs_time coordinate.
ds = trial_based_dataset(mode="stacked").drop_vars("trial_onset")
dsProblem 1: NDPointIndex requires matching number of variables and dimensions¶
NDPointIndex expects one coordinate variable per dimension. Our abs_time is a single 2D variable, not two 1D variables that define points in 2D space.
# This fails! NDPointIndex expects 2 variables for 2 dimensions
try:
ds.set_xindex(["abs_time"], xr.indexes.NDPointIndex)
except ValueError as e:
print(f"ValueError: {e}")ValueError: the number of variables 1 doesn't match the number of dimensions 2
What if we broadcast trial to 2D as well?¶
One might think: “NDPointIndex needs 2 variables for 2 dimensions. What if we broadcast trial to shape (trial, rel_time) to match abs_time?”
Let’s try this approach:
# Create 2D coordinates for NDPointIndex:
# 1. trial_2d - trial labels broadcast to 2D
# 2. event_locked - time relative to an event (different in each trial)
trial_labels = ds.trial.values # ['cosine', 'square', 'sawtooth']
trial_2d = np.broadcast_to(trial_labels[:, np.newaxis], (3, 500))
# Events happen at different times in each trial
event_times = np.array([2.0, 2.5, 1.5])
event_locked = ds.rel_time.values[np.newaxis, :] - event_times[:, np.newaxis]
ds_2d = ds.assign_coords(
trial_2d=(["trial", "rel_time"], trial_2d),
event_locked=(["trial", "rel_time"], event_locked),
)
ds_2d# Try to use NDPointIndex with trial_2d (strings) and event_locked
# This fails because NDPointIndex uses KDTree which requires numeric data!
try:
ds_2d.set_xindex(["trial_2d", "event_locked"], xr.indexes.NDPointIndex)
except ValueError as e:
print(f"ValueError: {e}")ValueError: could not convert string to float: np.str_('cosine')
Problem 2: NDPointIndex requires numeric coordinates¶
Because NDPointIndex uses a KDTree for spatial queries, all coordinates must be numeric. Our trial coordinate uses meaningful string labels like 'cosine', 'square', 'sawtooth' - a common pattern in experimental data. NDPointIndex cannot handle this.
To explore the other ergonomics issues with NDPointIndex, let’s create a version with numeric trial indices:
# Create numeric trial indices to work around the string limitation
trial_idx_2d = np.broadcast_to(np.arange(3)[:, np.newaxis], (3, 500))
ds_2d_numeric = ds.assign_coords(
trial_idx_2d=(["trial", "rel_time"], trial_idx_2d),
event_locked=(["trial", "rel_time"], event_locked),
)
# Now NDPointIndex can work with numeric coordinates
ds_ndpoint_2d = ds_2d_numeric.set_xindex(
["trial_idx_2d", "event_locked"], xr.indexes.NDPointIndex
)
ds_ndpoint_2dErgonomics Issue 1: Must provide ALL indexed coordinates¶
With NDPointIndex, you must provide values for all coordinates in the index. You cannot select by just one:
# With NDPointIndex, we MUST provide both coordinates
# This fails - can't select by event_locked alone:
try:
ds_ndpoint_2d.sel(event_locked=0.5, method="nearest")
except ValueError as e:
print(f"ValueError: {e}")ValueError: missing labels for coordinate(s): trial_idx_2d.
Ergonomics Issue 2: Point-wise queries, not dimensional selection¶
NDPointIndex supports querying multiple points using DataArrays, but these are point-wise queries - you specify exact (trial, event_locked) pairs. You cannot ask “give me all trials at event_locked=0.5”:
# Scalar query returns a single point
result_scalar = ds_ndpoint_2d.sel(trial_idx_2d=1, event_locked=0.5, method="nearest")
print("Scalar query (trial=1, event_locked=0.5):")
print(f" Shape: {dict(result_scalar.sizes)}")
# DataArray query returns multiple points - but you must specify EACH point explicitly
trial_query = xr.DataArray([0, 1, 2], dims="query")
event_query = xr.DataArray([0.5, 0.5, 0.5], dims="query") # Same event_locked for all
result_array = ds_ndpoint_2d.sel(
trial_idx_2d=trial_query, event_locked=event_query, method="nearest"
)
print("\nDataArray query (3 explicit points):")
print(f" Shape: {dict(result_array.sizes)}")
result_arrayScalar query (trial=1, event_locked=0.5):
Shape: {}
DataArray query (3 explicit points):
Shape: {'query': 3}
Ergonomics Issue 3: No slice support¶
NDPointIndex doesn’t support range queries with slice(). You can only query for single points:
# NDPointIndex doesn't support slices
try:
ds_ndpoint_2d.sel(trial_idx_2d=slice(0, 2), event_locked=slice(-0.5, 1.0))
except Exception as e:
print(f"{type(e).__name__}: {e}")ValueError: NDPointIndex only supports selection with method='nearest'
Ergonomics Issue 4: Semantic mismatch - “nearest in space” vs “value equals”¶
NDPointIndex uses Euclidean distance to find the nearest point. This creates unexpected behavior when the coordinates have different scales or meanings:
# Semantic mismatch: we ask for trial_idx_2d=0 and event_locked=1.0
# NDPointIndex uses Euclidean distance, so it might return a different trial
# if that's "closer" in the (trial_idx_2d, event_locked) space
result = ds_ndpoint_2d.sel(trial_idx_2d=0, event_locked=1.0, method="nearest")
result# Query for trial_idx_2d=1.5 (between trials) and event_locked=0
# NDPointIndex mixes trial and time to find the spatially nearest point
result2 = ds_ndpoint_2d.sel(trial_idx_2d=1.5, event_locked=0, method="nearest")
result2Comparison: How NDIndex handles the same data¶
Let’s see how NDIndex handles this dataset. We’ll also add an event_locked_time coordinate that represents time relative to an event that happened at a different time in each trial - this is the real use case for NDIndex:
from linked_indices import NDIndex
# Create fresh dataset with NDIndex instead of NDPointIndex
ds_ndindex = ds.assign_coords(
event_locked=(["trial", "rel_time"], event_locked)
).set_xindex(["abs_time", "event_locked"], NDIndex)
ds_ndindex# NDIndex: Select by event_locked time - "give me 0.5 seconds after the event in ALL trials"
# This is the key use case: same relative time across trials with different event times
result_event = ds_ndindex.sel(event_locked=0.5, method="nearest")
result_event# NDIndex: Slice by event_locked time - "give me -0.5 to +1.0 seconds around the event"
# Returns a bounding box containing all cells in that range across all trials
result_slice = ds_ndindex.sel(event_locked=slice(-0.5, 1.0))
result_sliceSummary: NDPointIndex vs NDIndex Ergonomics¶
| Aspect | NDPointIndex | NDIndex |
|---|---|---|
| Coordinate types | Numeric only (KDTree limitation) | Any type (strings, floats, etc.) |
| Select by single coord | Must provide ALL coords | Can select by any single coord |
| Query style | Point-wise (specify each point) | Dimensional (broadcast across dims) |
| Slice support | No | Yes (returns bounding box) |
| Query semantics | “Nearest point in N-D space” | “Cells where value matches” |
| Coordinate coupling | Coords are coupled (spatial distance) | Coords are independent |
The fundamental mismatch: NDPointIndex treats (trial_idx_2d, event_locked) as a 2D coordinate space and finds points by Euclidean distance. But for trial-based data, trial and time are independent dimensions - we want to select by event_locked alone without having to specify a trial.
NDIndex is designed for exactly this use case: selecting by values in N-D derived coordinates while preserving the dimensional structure of the data.
Summary¶
| Feature | NDPointIndex | NDIndex |
|---|---|---|
| Use case | Unstructured point clouds, curvilinear grids | Structured arrays with derived coordinates |
| Query type | Spatial: find nearest (x, y) point | Value: find cell where event_locked_time ≈ 7.5 |
| Coordinates | Multiple N-D coords (one per dimension) | Single N-D coord with computed values |
| Data structure | Points in N-D coordinate space | N-D array of scalar values |
| Slice support | No | Yes (bounding box) |